Common Problems
Once in a while, you may run into this problem,the experiment fails at atm either on 600 or 400. Do the following:
a). Stop the automation in IconNMR. Most of the time it will prompt you to change to another user before you can stop the automation. Follow the instructions on the pop-up window.
b). In Topspin window, type ii. Run ii a few times until you see no error messages. Then you can try atma and see if it works here(Better yet, you can manually tune and match the probe after typing in atmm.). If it finishes without errors, you can go back to iconnmr to restart the automation.
c). If running ii keeps on returning errors, you will need to restart Topspin.
On Fourier 300MHz, it will ask you to update the field once in a while. This is done under edlock's BSMS tap. On other Bruker spectrometers, if the field drifts so far off that you cannot lock the spectrometer, it calls for changing the base frequency. After that, expinstall all pulse sequences and parameter sets.
d). For nuclei other than 1H and 13C, especially those with large chemical shift ranges, it is important to set correct O1P and SW for regular 1D zg experiments to get good sensitivity to noise. Sometimes it is even necessary to run a few experiments for different chemical shift regions. The reason is that the excitation profile of a hard 90 degree (rectangular) pulse is gaussian shape. It falls to about 80% excitation at one half of 1/pw90 away from the spectral center. For example, for a pw90=10us, the excitation at +/- 50kHz from the offset is only about 80% of the excitation at the offset. You can use the shape tool in topspin to simulate the excitation profile. Type in stdisp in Topspin. This will bring you the shape tool window.
On Agilent 500MHz spectrometer,in the same study in walkup account you will need to run all 1D spectra before you run 2Ds, otherwise, a 1D and other followed experiments after a 2D will not run.
Common Questions
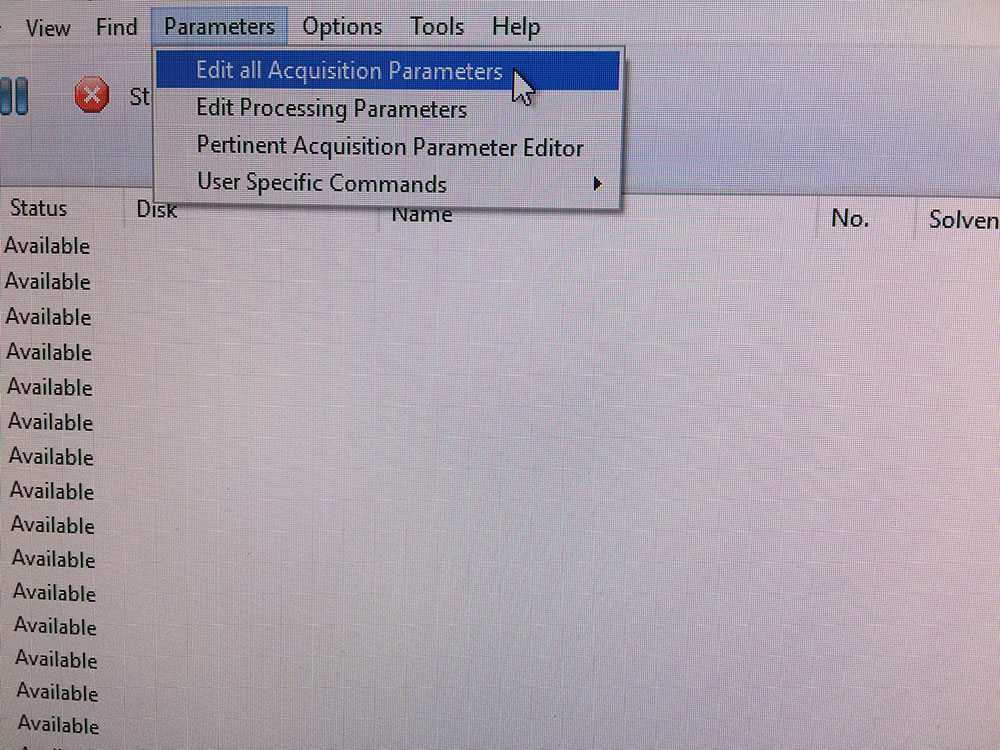
How to edit other acquisition parameters not shown in IconNMR?
Click on "Edit all Acquisition Parameters" to go to Topspin interface.
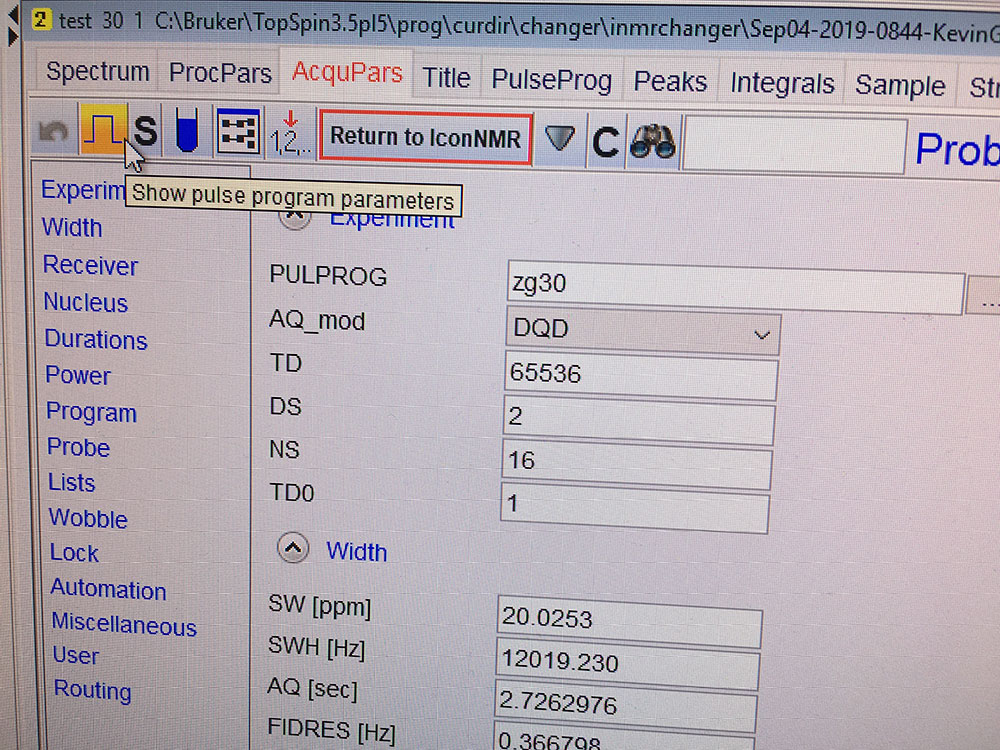
You can either edit parameters here or click on the square pulse shape button to go to another page which shows pulse program parameters only(recommended). After editing all necessary parameters, you'll need to click on "Return to IconNMR" to go back to IconNMR interface. DO NOT change the power levels unless you know what you are doing.
Topshim Options
The good starting shim called LASTBEST was generated by Xu in all Bruker spectrometers. DO NOT mess with that. You read the starting shim by typing in "rsh" in the command line, and choose LASTBEST. Then in the Topspin command line, type in "topshim gui". This will give you a Topshim window where you can choose options, and a shimming Report will be generated after the topshim is done. You only do Topshim 3D in water sample. And the default topshim shims up to Z5. You can choose to shim up to Z6 by selecting "use Z6". For even higher order Z(not recommended), you will need to type topshim ordmax=7 or 8 in the command line. It is always recommended to use "convcomp" option when sample is not spinning.